

The goal of vICC is to compute varying intraclass correlation coefficients (ICC) in a one-way random effects model (i.e., a random intercepts only model). Often computing an ICC is the first step when fitting a mixed-effects (a.k.a., hierarchical, multilevel, etc.) model that results in merely one value that is assumed to apply to each group (e.g., person, school, etc.). The underlying assumption is a common within-group variance, whereas, in vICC, a random-effects model is fitted to the residual variance, thereby permitting group-level ICCs. When subjects are the grouping variable, this is akin to investigating individual differences in the ICC.
The methodology in vICC was introduced in Williams, Martin, and Rast (2019). The context was measurement reliability in a cognitive task. To this end, vICC provides ICC(1), that is the correlation for any two observations from the same group, and ICC(2), that is average score reliability. Both ICC(1) and ICC(2) are reliability indices.
You can install the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("donaldRwilliams/vICC")vICC uses the popular Bayesian software JAGS to estimate the models. It must be downloaded from the following link: https://sourceforge.net/projects/mcmc-jags/files/
The following are implemented in vICC:
pick_group:
This model has a spike and slab on the random intercepts for the within-group variance. This provides posterior inclusion probabilities (PIP) that each group (e.g., person) does not belong to the common within-group variance model.
pick_tau:
This model has a spike and slab on the random effects standard deviation in the scale model which captures between-group variability in the within-group variances. This provides a PIP that there is variation in the within-group variances. In the context of reliability, a large PIP indicates that measurment invariance does not hold, given there are group-level differences in so-called measurment error.
pick_none:
This model also provides group-specific reliability, but there is no spike and slab formulation.
customary:
This is the standard random intercept model that assumes a common within-group variance.
Note that options 1 and 2 provide Bayesian model averaged estimates for the ICCs.
This is a basic example which shows you how to implement
pick_group
library(vICC)
library(ggplot2)
# congruent trials
congruent <- subset(flanker, cond == 0)
# subset 25 from each group
dat <- congruent[unlist(tapply(1:nrow(congruent),
congruent$id,
head, 25)), ]
# fit model
fit <- vicc(
y = dat$rt,
group = dat$id,
chains = 2,
iter = 500,
burnin = 10,
type = "pick_group"
)
fit
#> vICC: Varying Intraclass Correlation Coefficients
#> Type: pick_group
#> -----
#> Random Effects:
#> Post.mean Post.sd Cred.lb Cred.ub
#> RE.sd.mean 0.0467 0.0064 0.0362 0.0607
#> RE.sd.sigma 0.5241 0.0639 0.4056 0.6541
#> Cor(mean,sigma) 0.7929 0.0859 0.5983 0.9303
#>
#> Fixed Effects:
#> Post.mean Post.sd Cred.lb Cred.ub
#> FE.mean 0.4402 0.0070 0.4258 0.4529
#> FE.sigma 0.0954 0.0045 0.0869 0.1046
#> ICC(1) 0.1950 0.0471 0.1189 0.3022
#> ICC(2) 0.8511 0.0375 0.7714 0.9154
#> -----RE.sd.sigma is the between-group standard deviation for
the within-group variances, which is quite large and separated from zero
in these data. Further, Cor(mean, sigma) is the correlation
between the group-level means and the standard deviations.
The posterior inclusion probabilities are obtained with
pips <- pip(fit)
pips
Posterior Inclusion Probabilities:
#> Parameter Group PIP
#> RE_1 1 0.303
#> RE_2 2 0.283
#> RE_3 3 0.219
#> RE_4 4 0.998
#> RE_5 5 0.443
#> RE_6 6 0.346
#> RE_7 7 0.645
#> RE_8 8 0.323
#> RE_9 9 0.325
#> RE_10 10 0.268
#> RE_11 11 0.238
#> RE_12 12 1.000
#> RE_13 13 0.552
#> RE_14 14 0.339
#> RE_15 15 0.716
#> RE_16 16 0.913
#> RE_17 17 1.000
#> RE_18 18 0.992
#> RE_19 19 0.987
#> RE_20 20 0.998
#> RE_21 21 1.000
#> RE_22 22 0.997
#> RE_23 23 0.474
#> RE_24 24 0.398
#> RE_25 25 0.230
#> RE_26 26 0.427
#> RE_27 27 0.417
#> RE_28 28 0.945
#> RE_29 29 0.335
#> RE_30 30 0.255
#> RE_31 31 0.977
#> RE_32 32 0.330
#> RE_33 33 0.778
#> RE_34 34 1.000
#> RE_35 35 1.000
#> RE_36 36 0.227
#> RE_37 37 0.331
#> RE_38 38 0.348
#> RE_39 39 0.213
#> RE_40 40 0.249
#> RE_41 41 1.000
#> RE_42 42 0.999
#> RE_43 43 0.766
#> RE_44 44 1.000
#> RE_45 45 0.586
#> RE_46 46 0.852
#> RE_47 47 1.000
#>
#> ------which provide the probability that each group differs from the fixed effect within-group variance. The PIPs can then be plotted with
plot(pips)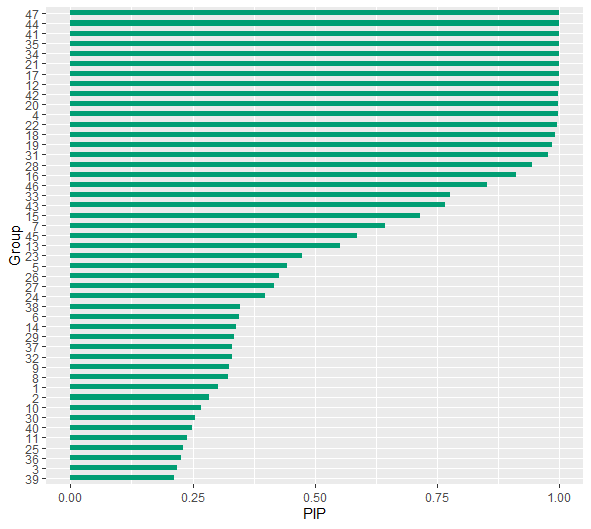
The group-level reliability, or ICCs, are plotted with
plts <- plot(fit)
plts$plot_icc1 +
theme(axis.text.x=element_text(angle=90, hjust=1),
legend.title = element_blank()) +
xlab("Group")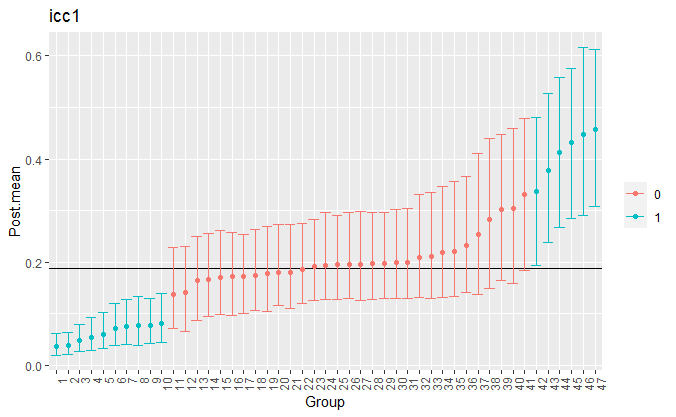
Notice that the object plts can be further modified with
ggplot2. Further, it also includes plots for the means
(plot_mean), standard deviations (plot_sd),
ICC(2) (plot_icc2).
Please cite Williams, Martin, and Rast (2019) when describing the
method and citation("vICC") when stating the software used
in the work.
Williams, Donald R, Stephen R Martin, and Philippe Rast. 2019. “Putting the Individual into Reliability: Bayesian Testing of Homogeneous Within-Person Variance in Hierarchical Models.” PsyArXiv.