
dplyrpins
DBI connectionsThe main goal of connections is to integrate
DBI-compliant packages with the RStudio IDE’s Connection
Pane. Packages such as RPostgres, RSQLite, RMariaDB and bigrquery connect R to
those databases, but do not provide a direct integration with the
Connections Pane. connections reads the configuration of
the connection and creates the integration with RStudio.
A second goal is to provide integration with the pins package. The
connections package allows you to pin database connections
and dplyr table objects.
Install the development version from GitHub with:
# install.packages("remotes")
remotes::install_github("rstudio/connections")The two main functions added by connections are:
connection_open() - Opens the database connection. Use
instead of dbConnect(), but use the exact same arguments.
It also automatically starts the Connections pane.connection_close() - Closes the database
connection.library(connections)
library(RSQLite)
con <- connection_open(SQLite(), "local.sqlite")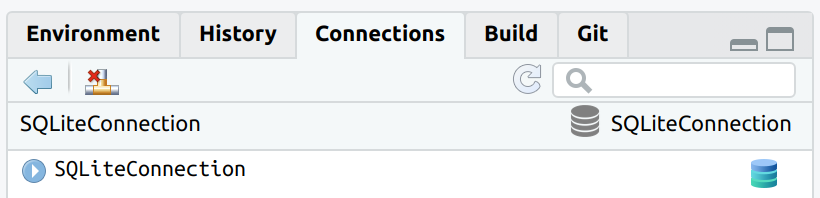
The connection can now be closed by using the appropriate button in
the Connections pane, or by using connection_close()
connection_close(con)
The connection code is parsed when connecting to the database, and it is visible once the connection is closed.
dplyrconnections integrates with dplyr by
supporting the following two functions:
tbl() - To create a pointer to a table or view within
the database.copy_to() - To copy data from the R session to the
database.The version of copy_to() inside connections
automatically updates the Connections pane, so the new table
automatically shows up.
con <- connection_open(SQLite(), "local.sqlite")
copy_to(con, mtcars, temporary = FALSE, overwrite = TRUE)
#> # Source: table<`mtcars`> [?? x 11]
#> # Database: sqlite 3.50.4 [/Users/edgar/Projects/connections/local.sqlite]
#> mpg cyl disp hp drat wt qsec vs am gear carb
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 21 6 160 110 3.9 2.62 16.5 0 1 4 4
#> 2 21 6 160 110 3.9 2.88 17.0 0 1 4 4
#> 3 22.8 4 108 93 3.85 2.32 18.6 1 1 4 1
#> 4 21.4 6 258 110 3.08 3.22 19.4 1 0 3 1
#> 5 18.7 8 360 175 3.15 3.44 17.0 0 0 3 2
#> 6 18.1 6 225 105 2.76 3.46 20.2 1 0 3 1
#> 7 14.3 8 360 245 3.21 3.57 15.8 0 0 3 4
#> 8 24.4 4 147. 62 3.69 3.19 20 1 0 4 2
#> 9 22.8 4 141. 95 3.92 3.15 22.9 1 0 4 2
#> 10 19.2 6 168. 123 3.92 3.44 18.3 1 0 4 4
#> # ℹ more rowsTo use an existing table inside the database use
tbl().
db_mtcars <- tbl(con, "mtcars")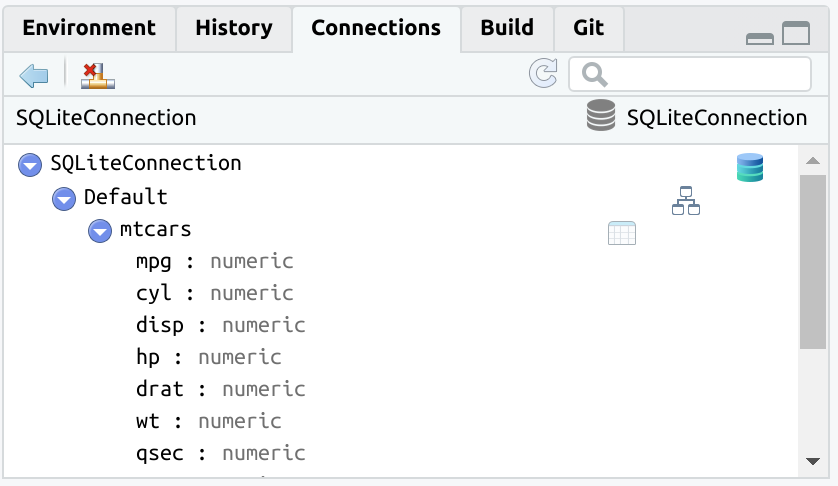
The tbl() function opens the rest of the already
available dplyr database integration.
db_mtcars %>%
group_by(am) %>%
summarise(avg_mpg = mean(mpg, na.rm = TRUE))
#> # Source: SQL [?? x 2]
#> # Database: sqlite 3.50.4 [/Users/edgar/Projects/connections/local.sqlite]
#> am avg_mpg
#> <dbl> <dbl>
#> 1 0 17.1
#> 2 1 24.4pinsThe connections package integrates with
pins. It adds the ability to “pin” database connections and
queries. It follows the same approach as the vetiver
package. connections now has two new functions:
connection_pin_write()connection_pin_read()The connection_pin_write() function does
not save the R object. It records the code necessary to
recreate the connection.
library(pins)
board <- board_folder("~/pins")
connection_pin_write(board, con, name = "my_conn")
#> Creating new version '20250910T182252Z-b2dcd'
#> Writing to pin 'my_conn'
If you wish to see the code that connections will use
when recreating the conneciton from the pin, you can use
connection_code():
connection_code(con)
#> library(connections)
#> library(RSQLite)
#> con <- connection_open(SQLite(), "local.sqlite")connection_pin_read() will replay the exact same code
used to initially connect to the database. Assign the output to a
variable, such as con1. The variable will work just like
any connection variable.
con1 <- connection_pin_read(board, "my_conn")The con1 variable is now a regular database connection
variable.
db_mtcars <- tbl(con1, "mtcars") %>%
group_by(am) %>%
summarise(avg_mpg = mean(mpg, na.rm = TRUE))
db_mtcars
#> # Source: SQL [?? x 2]
#> # Database: sqlite 3.50.4 [/Users/edgar/Projects/connections/local.sqlite]
#> am avg_mpg
#> <dbl> <dbl>
#> 1 0 17.1
#> 2 1 24.4dplyr database
queryWhen dplyr works with database data, the resulting query
is not executed until the data is explicitly collected into R, or when
printing the top results to the R Console. The pin records
two things:
The dplyr R object that contains all of the
transformations. It does not save the actual
results.
The necessary information to recreate the database connection. This is to make sure that the data is being retrieved from the original database connection.
connection_pin_write(board, db_mtcars, name = "avg_mpg")
#> Creating new version '20250910T182252Z-20f35'
#> Writing to pin 'avg_mpg'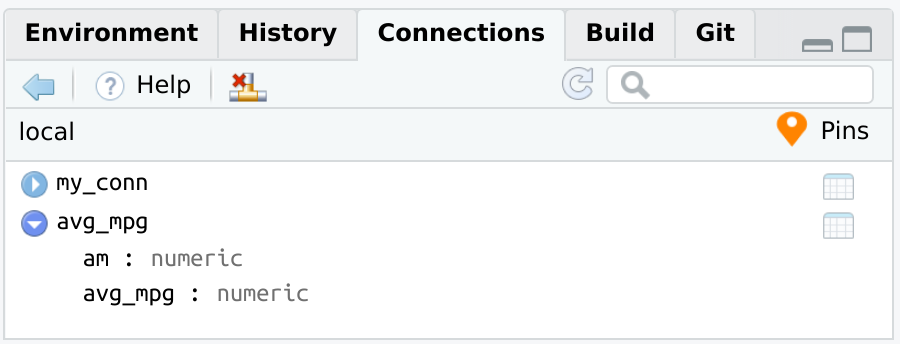
connection_pin_read() will connect to the database, and
return the dplyr object. Without assigning it to a
variable, the pin will immediately print the results of the database.
Those results are being processed at the time
connection_pin_read() runs.
connection_pin_read(board, "avg_mpg")
#> # Source: SQL [?? x 2]
#> # Database: sqlite 3.50.4 [/Users/edgar/Projects/connections/local.sqlite]
#> am avg_mpg
#> <dbl> <dbl>
#> 1 0 17.1
#> 2 1 24.4pins exampleThe way pins integrates with databases, via the
connections package, allows to open the connection from a
pin, and pipe all of the subsequent code into a new pin. Afterwards,
that pin can be used to collect or to continue using the
dplyr object.
board <- board_folder("~/pins")
con <- connection_pin_read(board, "my_conn")
tbl_summary <- con %>%
tbl("mtcars") %>%
group_by(cyl) %>%
summarise(avg_mpg = mean(mpg, na.rm = TRUE))
connection_pin_write(board, tbl_summary, name = "cyl_mpg")
#> Creating new version '20250910T182252Z-f8594'
#> Writing to pin 'cyl_mpg'
connection_close(con)
connection_pin_read(board, "cyl_mpg")
#> # Source: SQL [?? x 2]
#> # Database: sqlite 3.50.4 [/Users/edgar/Projects/connections/local.sqlite]
#> cyl avg_mpg
#> <dbl> <dbl>
#> 1 4 26.7
#> 2 6 19.7
#> 3 8 15.1
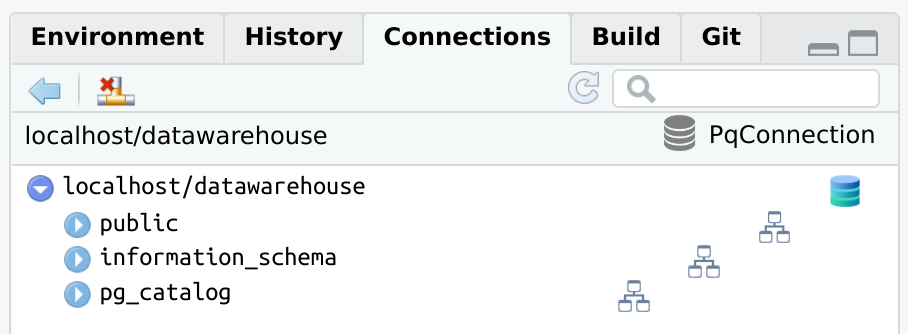
There are a couple of examples of how the Connections pane will look
when opening the connection via connections.
bigrquerylibrary(connections)
library(bigrquery)
con <- connection_open(
bigquery(),
project = "bigquery-public-data",
dataset = "austin_311",
billing = "my_project_billing",
use_legacy_sql = FALSE
)
connection_close(con)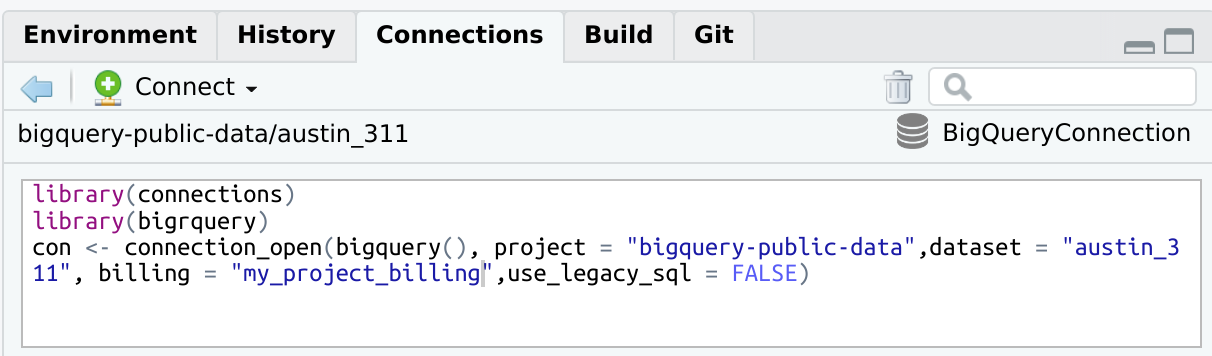
RPostgreslibrary(connections)
library(RPostgres)
con <- connection_open(
Postgres(),
host = "localhost",
dbname = "datawarehouse",
user = "[user id]",
password = "[password]",
bigint = "integer",
port = "5432"
)
DBI connectionsIt is possible to integrate DBI connections not opened
via connection_open(). To do that, use
connection_view() and pass it the variable containing the
existing database connection.
library(DBI)
con <- dbConnect(RSQLite::SQLite(), ":memory:")
connection_view(con)
Changes to the database will not automatically load in the
Connections pane. The connection_update() function will
refresh the pane with the latest.
dbWriteTable(con, "mtcars", mtcars)
connection_update(con)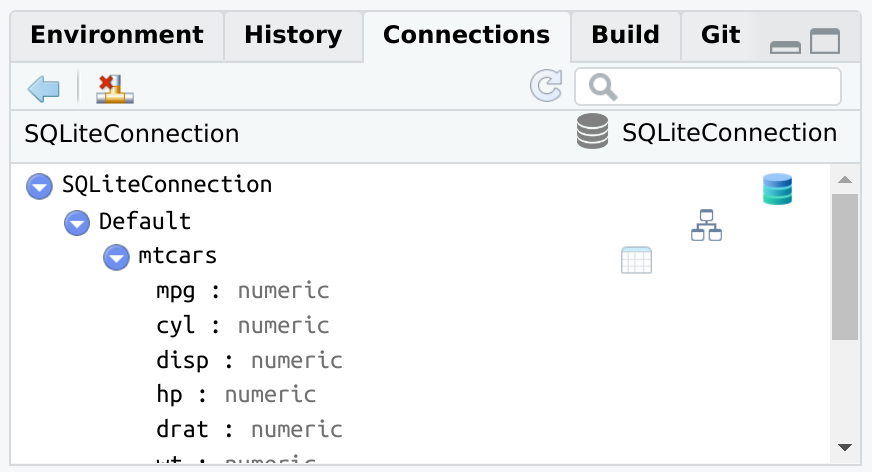
connection_close(con)